Though they live only a few hours before dividing, bacteria can anticipate the approach of cold weather and prepare for it. The discovery suggests that seasonal tracking is fundamental to life.

October 11, 2024
Every year, in latitudes far enough north or south, a huge swath of life on Earth senses that winter is coming. Leaves fall from trees, sparrows fly to the tropics, raccoons grow thick winter coats, and we unpack our sweaters from storage. Now scientists have shown that this ability to anticipate shorter days and colder temperatures is more fundamental to life than anyone thought: Even short-lived, single-celled organisms can sense day length and get themselves ready for winter.
Lab experiments, recently published in Science, show that cyanobacteria — a type of bacteria that produces energy from sunlight through photosynthesis — anticipate the change(opens a new tab) by bundling up in their own way. They turn on a set of seasonal genes, including some that adjust the molecular composition of their cell membranes, to improve their odds of survival.
The study authors were amazed to find this season-sensing ability in an organism that lives for only about five hours in the lab before dividing. “It seemed like a very nonsensical idea to think that bacteria would care about something that’s happening on a scale that’s so much bigger than their lifetime,” said Luísa Jabbur(opens a new tab), a microbial chronobiologist at the John Innes Center in Norwich, England, and lead author of the new paper.
But cyanobacteria have an evolutionary incentive to pass on relevant information to their progeny: Each cell divides into two identical clones, and each of those does as well, ad infinitum. Carl Johnson(opens a new tab), the senior paper author at Vanderbilt University, likened it to the way monarch butterflies migrate south for the winter but never make the return journey north — their offspring do that. “When you start thinking about more of a lineage, or as the colony or population,” he said, “then that kind of thing makes perfect sense.”
The discovery connects cyanobacteria to a plethora of much more complex organisms with seasonal rhythms, and it indicates that anticipating seasons may have emerged early in life’s evolution. It may have even predated the internal clocks that give an organism a sense of day and night. “This issue of dealing with seasonality may be very fundamental to why [biological] clocks exist in the first place,” said the cell biologist Mike Rust(opens a new tab), who studies cyanobacteria’s internal rhythms at the University of Chicago and was not involved in the new research. Staying in sync with the seasons could be more ancient and more elemental to life than anyone suspected.
How Cells Keep Time

When Johnson entered graduate school in the 1970s, scientists knew that circadian clocks — organisms’ internal timekeepers for the day-night cycle — are ubiquitous in multicellular plants and animals. These molecular devices choreograph delicate dances, such as plants unfolding their leaves in the morning and closing them at night. (They’re also the reason why humans have definitive sleeping and waking hours, as well as disjointed sensations when traveling between time zones or pulling an all-nighter.)
But the idea that simple organisms such as bacteria could have daily clocks as well was deemed controversial. Johnson looked into the possibility in graduate school, to no avail. Then, in 1986, evidence emerged that cyanobacteria do indeed have daily rhythms. When the South African plant physiologist Nathanaël Grobbelaar exposed cyanobacteria to light and dark periods, he observed that the cells processed nitrogen, a key nutrient, only during the simulated night(opens a new tab). It was the first record of a day-night internal rhythm in any single-celled organism.
The discovery gave Johnson an idea: If cyanobacteria have daily rhythms, maybe he could identify the molecules that, like gears in a watch, make the organisms’ circadian clock run. In papers published in 1993(opens a new tab) and 1998(opens a new tab), with collaborators in Japan and Texas, he identified three genes and their corresponding proteins — KaiA, KaiB and KaiC (kai is Japanese for “cycle”) — involved with the cyanobacterial circadian clock. Interactions between KaiA and KaiB create a reaction in which KaiC acquires an extra phosphate group and then sheds it rhythmically, in sync with day and night. Astonishingly, the scientists also found that the whole loop can happen outside a cell, among loose molecules in a test tube.
Cyanobacterial colonies pulse through a day-night cycle as a gene involved in the circadian clock cycles on and off. Biologists attached a bioluminescent reporter gene to the clock gene to visualize the rhythm of the cells’ circadian clocks. The brighter color indicates higher expression of the clock gene.
Courtesy of Carl Johnson
Since then, researchers have learned a lot about the cell biology underlying these rhythms. But it would take another quarter of a century to connect those same genes to an ability stretching over a longer time frame, one that is more calendar than clock.
Winter Is Coming
One day in 2018, as Jabbur scoured the literature on cyanobacteria’s circadian clock, she realized something was missing. She could not find any explorations of the relationship between the circadian clock, which follows Earth’s axial, day-night rotation, and a seasonal rhythm linked to how the Earth’s axis is tilted, wherein summer happens in a hemisphere that tilts toward the sun.
“That was a bit shocking,” Jabbur recalled, because cyanobacteria’s circadian clock is the best studied of any organism. She wondered whether the same proteins might lead to what’s known as a photoperiodic response — the ability to react to the length of a day and “use the information to change their physiology, metabolism or behavior in anticipation of upcoming seasons,” she said.
She brought the idea to Johnson, her doctoral adviser. Initially he laughed at it. Cyanobacteria make food from light, so it seemed obvious that the cells would thrive during longer days and suffer when the periods of light grew shorter. But he told Jabbur to try the experiment anyway because, as a sticky note on his office door states, “progress is made by young scientists who carry out experiments that old scientists say would not work.”

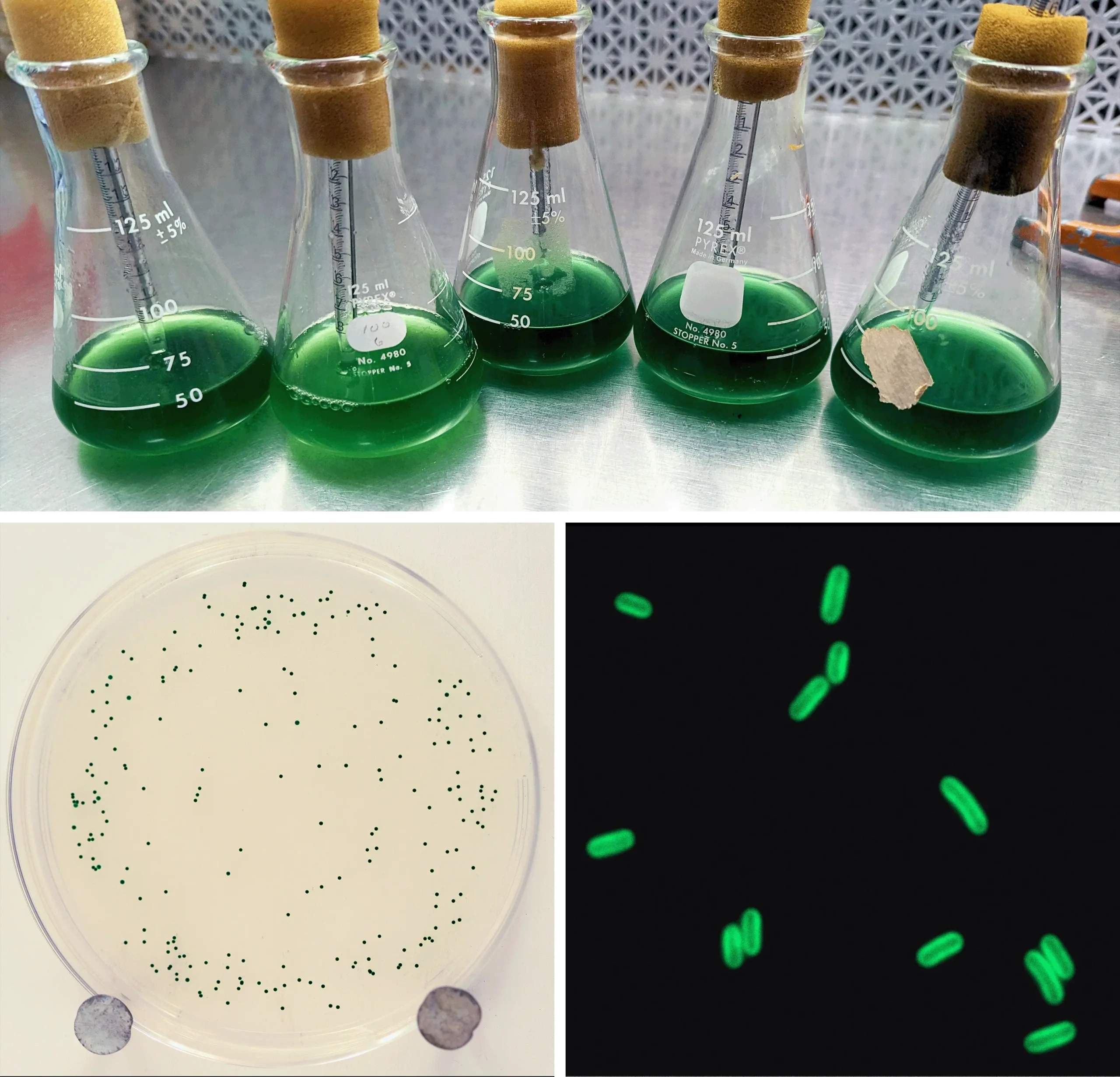
She proved her mentor wrong, and the note right, almost immediately. Within a week, she appeared in Johnson’s office with two bacterial plates. Both had been plunged into ice water to simulate the onset of winter. But one hosted more visibly green cyanobacteria than the other. The bacteria that thrived had been exposed to longer dark periods beforehand — they’d had an opportunity to anticipate what was coming.
In an expanded experimental protocol, reported in the new Science paper, Jabbur exposed three groups of cyanobacteria to different periods of light and darkness for eight days, representing winter (eight hours of light and 16 hours of darkness), equinox (12 hours of light and 12 hours of darkness) or summer (16 hours of light and eight hours of darkness). Then she dunked them in the ice water, sampled bacteria from each chilled tube and watched for colonies to grow from live cells.
Despite growing at warm temperatures, the cells that experienced short, winterlike light durations seemed to know that cold was coming and were able to prepare for it. They survived up to three times better after the frigid bath than either the summer or equinox cells. But how?
Jabbur compared the genes activated in the different groups of cells. The winter-condition cells expressed more genes related to metabolism, while the summer-condition cells expressed genes related to heat and ultraviolet light, suggesting that they had adapted for a different season. She looked at one change more closely: the molecular composition of their cell membranes.
It is well known that cell membranes, including those that encircle cyanobacteria, are sensitive to temperature. Like butter, the lipids that compose membranes become more rigid in cold conditions and more fluid in heat. Many organisms can adjust their membranes — a process known as desaturation — to keep molecules moving freely across the membrane in a range of temperatures. Jabbur wondered if her cyanobacteria were doing the same thing. Indeed, further experiments showed that her winter-primed cyanobacteria had more desaturated lipids that kept their cell membranes from gumming up as the temperature dropped.
Finally, she wanted to know if these photoperiodic adaptations were tied to the circadian clock or driven by a separate mechanism. When the researchers deleted the genes that encode the KaiA, KaiB and KaiC proteins, the winter-condition cells survived no better than summer-condition cells. They had failed to adjust their lipids. The daily molecular clock might be driving the seasonal calendar as well.
“We still don’t know if the clock is the one that is actually encoding the day length,” Jabbur said. “But it appears to be necessary for the response.”
An Ancient Talent
Cyanobacteria are the most ancient known life form that still lives on Earth, encompassing billions of years of history. About 2.4 billion years ago, they transformed the chemistry of our atmosphere to the oxygen-rich mix we enjoy today. It is humbling to think that something so ancient and small may contain the seeds of the complex seasonal anticipation behaviors we see today, from the migration of shorebirds and songbirds, to hibernating grizzly bears, to the human craving for pumpkin spice lattes.
“It is truthfully impressive that organisms as old as cyanobacteria could have this kind of response,” Jabbur said. “It makes one really wonder about when [photoperiodism] first emerged, and what Earth looked like back then.”
Because organisms go through daily cycles more frequently than seasonal ones, scientists have generally assumed that circadian clocks evolved before photoperiodism. But the new research suggests another possibility. “Photoperiodic measurement could have been the first thing [to evolve],” Johnson said. Perhaps our oldest ancestors needed to invent an internal clock to survive the stresses of seasonal weather — and then daily cycles were built on top of that.
What remains puzzling, however, is how such a short-lived organism could evolve a mechanism to track time through entire seasons, which stretch hundreds of times longer than its own lifetime. “One intriguing issue is whether or how these signals are passed on through cell generations, since seasonal changes happen much more slowly compared to the generation time of these cells,” said Devaki Bhaya(opens a new tab), a senior staff scientist at Carnegie Science who was not involved in the research. No matter how it happens, the mechanisms wouldn’t have been selected for individual survival, but for the welfare of the entire genetic line, encompassing many generations of cyanobacteria.
Still, these ideas remain speculative as long as photoperiodism is identified in only a single species of cyanobacteria. In her new role as a research fellow at the John Innes Center, Jabbur plans to explore the photoperiodic responses of more bacteria to better understand when this ability to anticipate seasons might have evolved. Other strains of bacteria have circadian clock genes that drive mechanisms markedly different from those of cyanobacteria. They may reveal more secrets about internal rhythms and seasonal adaptations. Only time will tell.











